Mercator Fellowships
Dr. Jiri Jablonsky
Multi-omics data integration into mathematical model to analyze the metabolic regulation of cyanobacteria
My group is focusing on metabolic regulation of cyanobacteria, including the local and global roles of isozymes. The computational method I have developed is integrating various omics and growth data to analyse shifts between different growth conditions and regulatory switches. My previous work determined the essential role of isozymes in high and ambient CO2 growth conditions, suggesting a similar role of isozymes also for autotrophy-heterotrophy switch. Furthermore, we plan to do in silico experiments to decipher the regulation of triose phosphates and isozymes within glycogen metabolism, and to further analyse Ci acclimation.
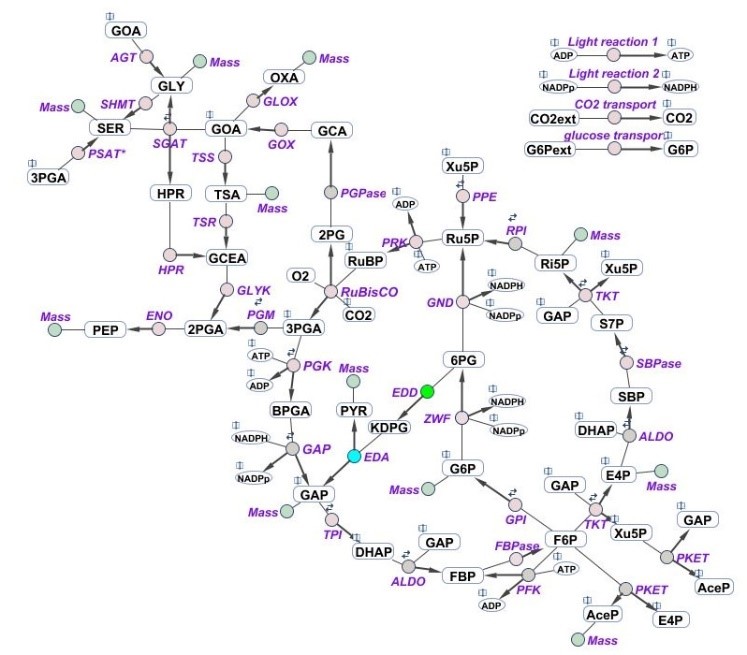
Schematic representation of the central carbon metabolism network implemented in the multi-level kinetic model of Synechocystis sp. PCC 6803.
The model consists of Calvin-Benson cycle, photorespiration, all four glycolytic pathways and simplified carbohydrates synthesis and biomass production. TCA cycle will be added during the project.
Prof. Conrad Mullineaux, Mercator Fellow 2018 - 2021
Fluorescence microscopy to probe distributions, dynamics and interactions of cell components in cyanobacteria
My group works on the cell biology of cyanobacteria, and we have developed expertise in using fluorescence microscopy to visualise and quantify the sub-cellular distribution and dynamics of cell components including proteins, nucleic acids and fluorescent metabolite analogs. For this programme we are helping members of the SCyCode team to apply these approaches to understand the regulation of metabolism at the single-cell and sub-cellular scales in Synechocystis.
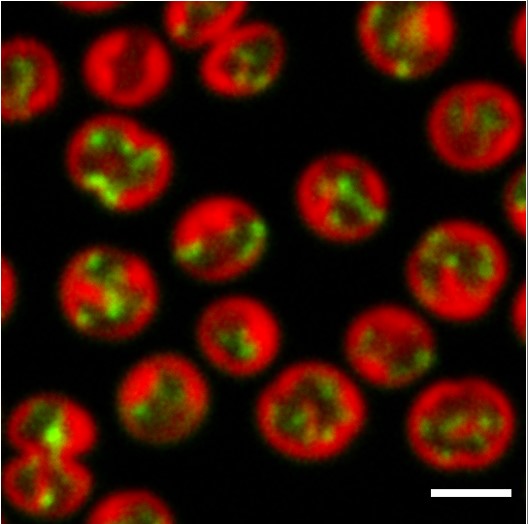
Fluorescence microscopy displays the localization of psaA mRNA (in green) and thylakoid membranes (in red) in Synechocystis cells.
Principal Investigator

University of South Bohemia
Institute of Complex Systems
Czech Republic
jiri.jablonskygmail.com
Principal Investigator

Queen Mary University of London
United Kingdom
c.mullineauxqmul.ac.uk